Just noticed this while working and debugging covid5ync app. Don't know if that may have sense from a biotechnology view... still, it worth to be mentioned:
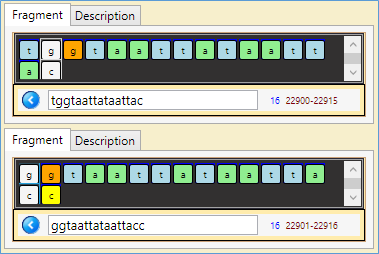
- A fragment (15 nucleotides) of the RNA sequence looks to be a 'palindrome' -:) of itself… Position 22901 -22916
Note: the top part of the figures below reads sequence's nucleotides in the direction 5'->3', the lower should be read in the reverse direction 3'->5' (which is the complementary of the upper part (i.e. 'a' complementary to 't' and 'g' to 'c'))…
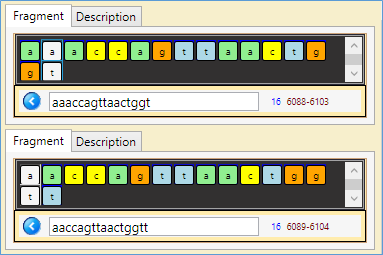
- A similar 'palindrome' fragment (15 nucleotides): 6089 - 6104
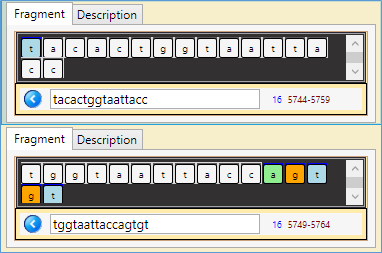
- Similar with a small shift: (5744 – 5764)
I continue posting about covid-5ync app (a helper app contribution in the fight against the latest coronavirus epidemic)
A major step now done: serializing application's information into xml files!
Hopefully this will allow transmitting research session work among members of our target community: biotechnology engineers.
I also downloaded (@NCBI site) a recent version of the virus RNA sequence (referred to as: MT163719 29903bp RNA linear VRL 10-MAR-2020). A daily work is done by the application to retrieve significant regions on the sequence. The updated version of the (xml) file is then uploaded to the project's web site which will allow gaining time in some tedious analysis tasks.
Serialization / deserialization
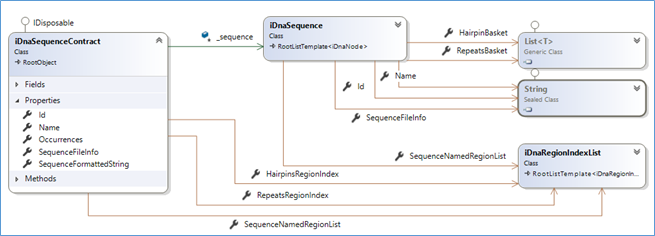
Reminder: Technically speaking, the sequence's data structures can be summarized as:
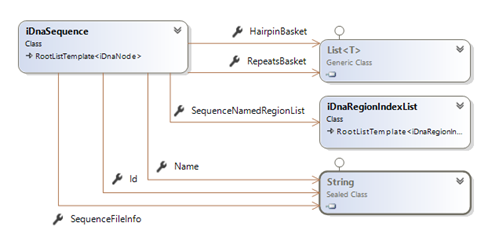
To serialize these information… (in regards of the urgency matters) I decided to simply use a DataContractSerializer (which, as I said in other posts, is not really the best solution for extensibility).
The encountered difficulty was on several aspects:
- Some information is redundant, notably for sub-sequences (collections of repeats and hairpins)
- The sequence object being itself a collection of nodes, the serializer did not really allow an easy way to handle its sub-objects. Here we fall into a known problem of cycling!
The solution used was:
- Transform sub-sequences to region lists (start/end indexes + name + occurrences) and deserialize back to sub-sequences
- Create a serialization-specialized object (using DataContract / DataMember attributes) and process the serialization through that object to avoid the 'cycling' errors.
The second difficulty was errors encountered on deserialization while locating and assigning various regions start/end indexes to the sequence's nodes.
Actually, the deserialization submits the sequence's node as a string to be parsed asynchrony. And as you may have guessed, the collection of sequence's nodes was being altered while locating the regions' nodes!
The solution:
- The Parse method raises Parse Complete event
- Subscribe to that event and proceed to these operations.
That is the short story… will keep you posted about more details later this week!
KEEP SAFE: wash your hands + do not touch your face + keep hope: Humanity will prevail!
This post, follows in the series about technical details of covid-5ync (an open source application contribution in the fight against the new covid-19 virus).
This time, our talk is about searching nucleotides within a dna sequence.
Search is an important task for analyzing a sequence. Actually, to understand the structure of a genome, one task is to locate and classify identical regions and complementary regions (complementary regions = where nucleotides are paired: aót, góc). From the IT point of view, such a task is obviously related to 'search'… and should particularly be optimized.
Using Linq
To locate a sequence of nucleotides (i.e. a string) within a (global) sequence, we may simply iterate into the global sequence nodes (each of which is a char) until we find the searched string. To find all occurrences, we can repeat the process starting at the next location and so on.
Despite the artisanal aspect of the process, that works well for searching a predefined string.
Another task that looks related to search, is to find 'repeats' (Repeats are identical regions on a sequence).
It is somehow different from searching a predefined string. The application actually has to iterate through the sequence, and at each position take a number of nucleotides to compose a string to be searched all over the sequence… (and keep coordinates of the found occurrences).
Proposed solution
In covid-5ync, a dna sequence is a List<dna node> where each node contains an Index (int). Using Linq, we can define a method that returns the string of a given length at a given location (node Index) of the sequence:
public string StringAtIndex(int index, int len)
{
string str = "";
var nodes = this.SkipWhile(i => i.Index < index).Take(len);
foreach(var n in nodes)
str += n.Code.ToString();
return str;
}
|
With this method in place, we can efficiently locate all starting nodes of occurrences of a string on the sequence:
IEnumerable<iDnaNode> AllStartoccurrencesOfString(string str)
{
int len = str.Length;
return this.Where(i => StringAtIndex(i.Index, len) == str);
}
|
Sample usage: locate and select occurrences of a string
// get all starting nodes of the string
Var allStarts = AllStartoccurrencesOfString(str).ToList();
// visit the found starting nodes and select (str.Length) consecutive nodes…
foreach( var item in allStarts)
{
var subSeq = this.SkipWhile(n => n.Index < item.Index).Take(str.Length);
subSeq.SelectAllNodes();
}
|

A quick post to talk about the progress in this project with a few technical details.
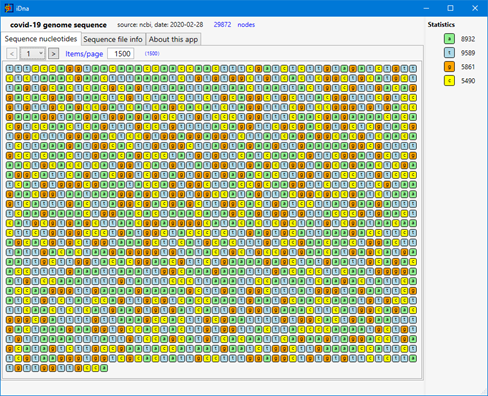
First, a screen capture of what it looks like today (just a little better than before!):
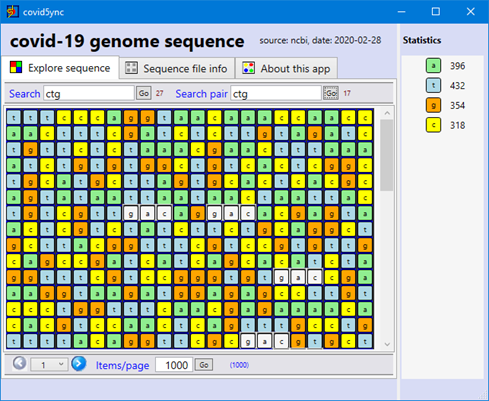
The first version of the application is already online: click-once installation. And Yes, we don't have money to have a certificate for the deployment… install only if you trust!... any suggestions about sponsors are also greatly welcome!
A few features implemented:
- Search for string and search for the 'complementary' of string.
- Paging
- Zoom-in / out on the sequence
Next steps:
- Search enhancement: visually locate occurrences
- Search for unique fragments according to user-provided settings
- Open file / save selection
Long-term:
Now, let us take a quick dive into the mechanics
The first class diagram (updated source code with such documentation is @github)
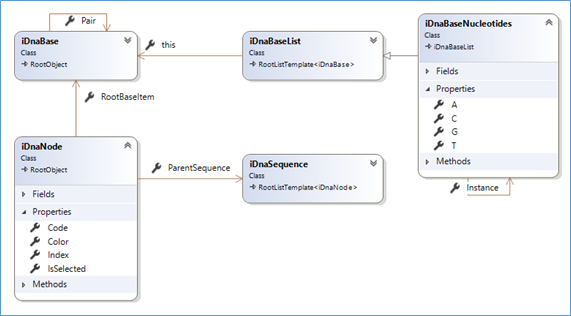
That seems quite basic(!), actually, as the reality of DNA, and that is one reason it is also fascinating!
- A sequence (iDnaSequence) is composed of 'nodes' (iDnaNode)
- A node is noted as a char. It refers to a 'base'.
- Commonly a set of 4 bases is used ('a', 't', 'g' and 'c'). There may be more, but we can easily handle this when needed.
- Each individual base has a complementary 'Pair'. A is pairable with T, C with G
- The 'standard' set of bases (iDnaBaseNucleotides) is a (singleton) list of the 4 bases. It sits as the main reference for nodes. It provides answers to important questions like: is 'c' a valid base?... what the complementary of 'X'?... what is the complementary sequence of the string "atgccca"? and so on.
Visual presentation: a start point
There are many ways to present a DNA sequence. To start with something, let us assign a color to each base. The user can later change this to obtain a view according to his or her work area. Technically speaking, we have some constraints:
- The number of nucleotides of a sequence can be important. For coronavirus, that is roughly 29000. We therefor need 'paging' to display and interact with a sequence.
- Using identification colors for nucleotides can also help to visually identify meaningful regions of the sequence on hand. For this to be useful, we need to implement zoom-in/out on the displayed sequence.
Paging
I simply used the solution exposed in a previous post about doc5ync.
Zoom-in / out
I found a good solution through a discussion on Stack Overflow
(credit: https://stackoverflow.com/users/967254/almulo).
- Find the scroll viewer of the ListView.
- Handle zoom events as required
var presenter = UiHelpers.FindChild<ScrollContentPresenter>(listItems, null);
var mouseWheelZoom = new MouseWheelZoom(presenter);
PreviewMouseWheel += mouseWheelZoom.Zoom;
|
Sample screen shots of a zoom-out / in

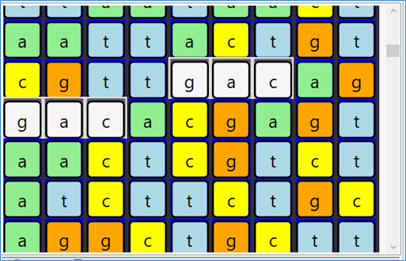
More details later in a future post.
Please send your remarks, suggestions and contributions on github.
Hope all that will be useful in some way… Time is running… Humanity will prevail
NCBI (National Center for Biotechnology Information) offers a vast database of DNA sequences.
I visited their site to see if I can find information about the last version of the menacing coronavirus.
Yes, that is available. It is precisely named: coronavirus 2 (SARS-CoV-2), and a long list of its DNA sequences is there.
I had worked, long years ago, on DNA sequences analysis and felt like giving a try to see how that sequence can be presented… just to see!
My old app (MFC, C++ app) could open and analyze the sequence. DNA sequence analysis is a long story. As far as I could learn: locate repeats (fragments that are repeated on the sequence), locate 'hairpins' (fragments of complementary nucleotides: a<=>T and g<=>c)… etc.
The old app did not seem quite handy to manipulate the downloaded sequence, so I started writing a new WPF one.
A few hours later, the app could display the sequence in a somehow 'visual appealing' UI, which invited to go ahead for some more significant work.
Covid-19 is not for fun!
Yes, it is not really for fun! I am not yet sure how such work can be useful, but whatever effort everyone can provide might be of help in defeating this new danger. Let us start and see!
For now, what I intend to do is:
- Port the biotechnology features of the old app to a new handy UI;
- Publish the app online for biotechnology engineers working on the subject: and get their feedback
- Upload the source code to github for IT community feedback and contributions
It is a very small step in a long way to defeat that epidemic.
More on this in the next few days / weeks.
Be safe!